
Longitudinal Bout Detection
Source:vignettes/longitudinal_bout_detection.Rmd
longitudinal_bout_detection.RmdIntroduction
This vignette demonstrates how to detect and analyze song bouts across longitudinal recordings using SAP objects. Bout detection identifies continuous periods of vocalization, which is essential for understanding song production patterns during development.
Prerequisites: Before reading this vignette, we recommend completing:
- Overview: ASAP 101 - Basic ASAP functions
- Constructing SAP Object - SAP object creation
- Longitudinal Motif Detection - Motif-based workflow
Overview
What are bouts? Song bouts are continuous periods of vocalization separated by silence. In zebra finches, a bout typically contains multiple motif renditions and provides important context for understanding song structure and production.
Why detect bouts? Bout-level analysis enables:
- Tracking song production rate across development
- Analyzing bout duration and structure changes
- Understanding motif density within bouts
- Identifying practice patterns and song maturation
Relationship to motif detection: While motif
detection identifies specific, repeatable vocal sequences, bout
detection captures the broader temporal structure of vocalization. When
used together (with summary = TRUE), you can analyze how
motifs are organized within bouts across developmental time points.
Complete Pipeline
The bout detection workflow processes all recordings in a SAP object to identify bouts across multiple time points.
Option 1: Load SAP Object from Previous Analysis
If you’ve already completed motif detection (see Longitudinal Motif Detection), you can load that SAP object and add bout detection:
library(ASAP)
# Load SAP object from previous motif detection vignette
sap <- readRDS("longitudinal_motif_analysis.rds")
# Add bout detection with motif-bout relationships
sap <- sap |>
find_bout(
rms_threshold = 0.1,
min_duration = 0.4,
gap_duration = 0.3,
freq_range = c(3, 5),
summary = TRUE # Include motif-bout relationships
)
# Visualize results
plot_heatmap(sap, segment_type = "bouts", balanced = TRUE)Option 2: Create New SAP Object
Alternatively, you can create a new SAP object from scratch (see Constructing SAP Object for detailed instructions):
library(ASAP)
# Create SAP object
sap <- create_sap_object(
base_path = "/path/to/recordings",
subfolders_to_include = c("190", "201", "203"),
labels = c("BL", "Post", "Rec")
)
# Detect bouts across all recordings
sap <- sap |>
find_bout(
rms_threshold = 0.1,
min_duration = 0.4,
gap_duration = 0.3,
freq_range = c(3, 5)
)Note: When creating a new SAP object without prior
motif detection, summary = TRUE will not add motif-bout
relationship columns (n_motifs, align_time, etc.). Additionally, because
template matching is not performed, it is possible to detect false
positives such as innate vocalizations or background noise.
Understanding SAP Object Behavior
The find_bout() function processes recordings based on
the SAP object’s metadata:
- Lazy loading: Audio files are read on-demand during processing
-
Centralized storage: Results are stored in
sap$bouts -
Optional visualizations: Set
save_plot = TRUEto save detection plots - Parallel processing: Uses multiple cores for efficient batch processing
Example Output
# View bout detection summary
summary(sap$bouts)
# Example output:
# filename day_post_hatch label selec start_time end_time n_motifs
# S237_190_1.wav 190 BL 1 2.45 8.32 12
# S237_190_1.wav 190 BL 2 15.67 19.84 8
# S237_201_1.wav 201 Post 1 3.21 10.45 15When summary = TRUE is enabled with existing motif data,
additional columns are included:
-
n_motifs: Number of motifs detected within each bout -
align_time: Time of first motif (useful for alignment) -
bout_number_day: Sequential bout number within each day -
bout_gap: Time interval from previous bout (within same file)
(Optional) Saving SAP Object
After completing bout detection, you can save the SAP object for later use:
# Save the SAP object with bout detection results
saveRDS(sap, "longitudinal_bout_analysis.rds")
# Later, load it back
sap <- readRDS("longitudinal_bout_analysis.rds")What gets saved: - All metadata and file references - Bout detection results - Motif data (if previously detected) - All spectral features and analysis results
Important notes: - The original WAV files are
not included in the saved object - You must keep the
WAV files at their original paths to run additional analyses - The saved
.rds file is typically much smaller than the audio data
Visualizing Results
Bout Spectrograms
Visualize sample bouts from each time point to verify detection quality:
# Visualize 3 random bouts per time point
visualize_segments(sap, segment_type = "bouts", n_samples = 3, by_column = TRUE)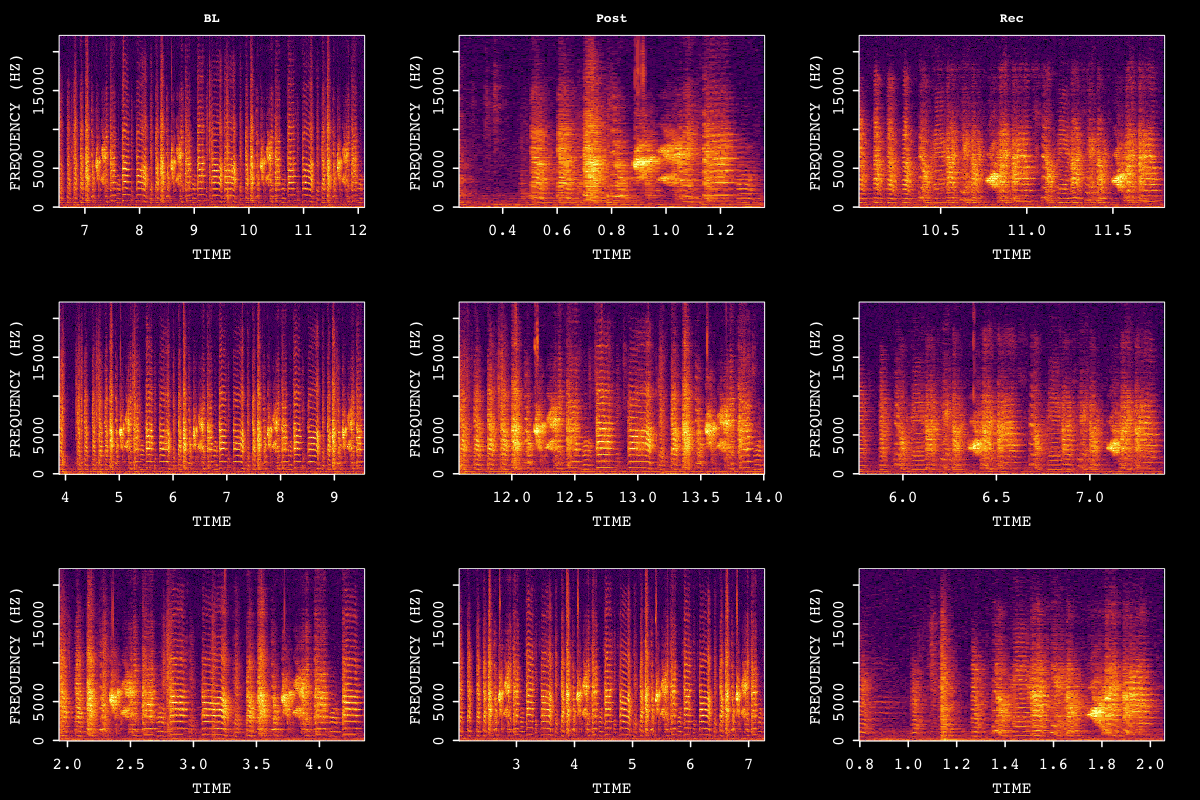
Amplitude Envelope Heatmap
Amplitude envelope heatmaps visualize the temporal structure of detected bouts across song developmental stages.
# Create amplitude envelope heatmap with balanced sampling
plot_heatmap(sap, segment_type = "bouts", balanced = TRUE)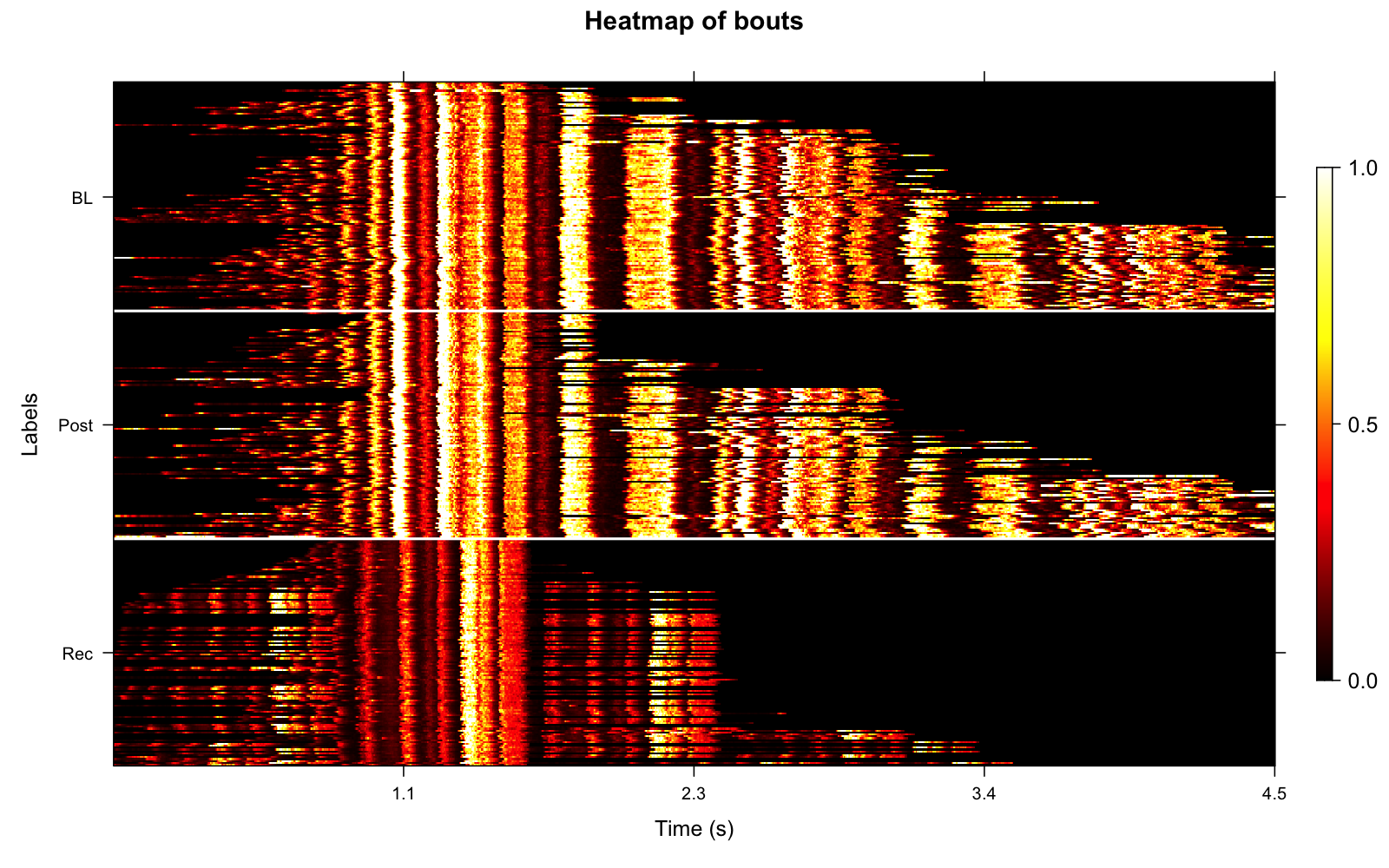
The balanced argument ensures equal
representation across time points (e.g., same number of bouts from each
developmental stage), which improves statistical comparisons of bout
structure across development.
Bout Summary Statistics
Understanding Summary Metrics
When summary = TRUE is enabled and motif data exists,
find_bout() calculates additional metrics that reveal the
relationship between bouts and motifs.
Example bout summary table (showing first 10 bouts):
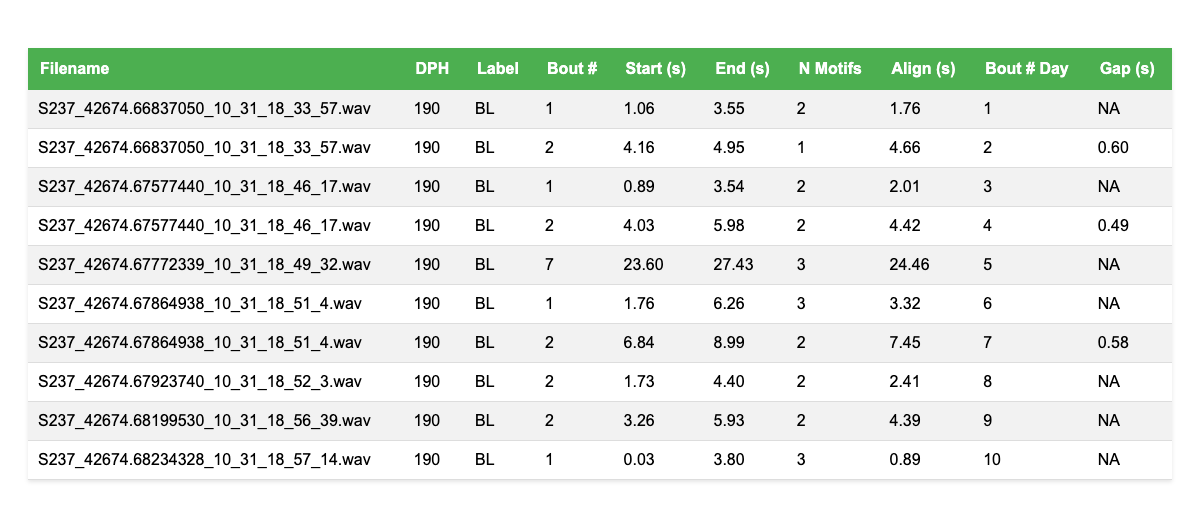
The table includes all detected bouts with the following columns:
-
n_motifs: Count of motifs within each bout- Reveals motif density and bout complexity
- Typically increases during song development
-
align_time: Timestamp of first motif in bout- Useful for aligning bouts across recordings
- Enables time-locked analysis of bout structure
-
bout_number_day: Sequential bout number within each day- Tracks bout order within recording sessions
- Useful for analyzing practice patterns
-
bout_gap: Time interval from previous bout- Measures inter-bout intervals
- Reveals temporal patterns in song production
Interpreting Developmental Changes
Bout statistics can reveal important developmental patterns. The plots below show three key bout metrics across developmental stages:
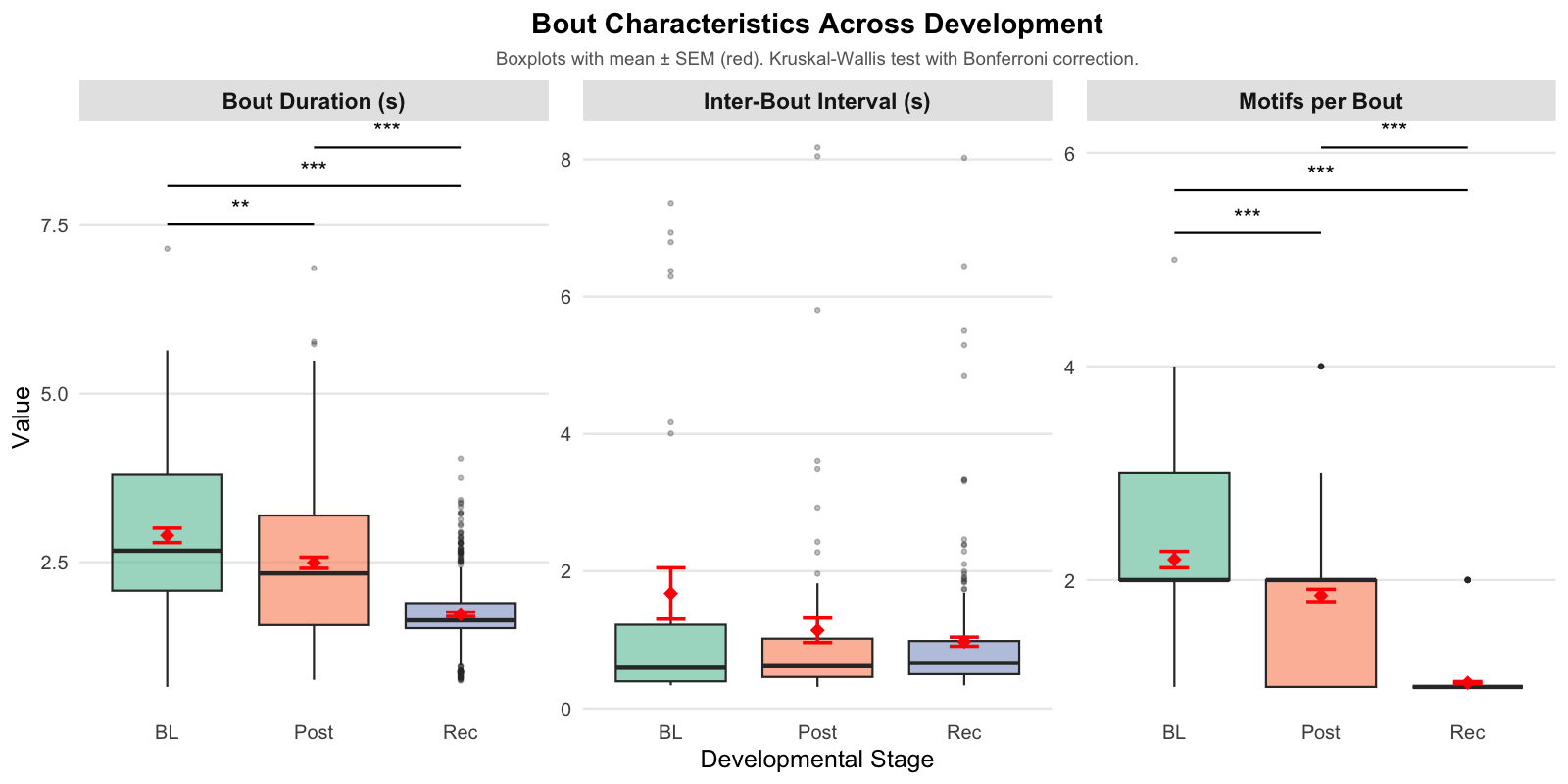
Understanding the visualization:
-
Boxplots (colored boxes and whiskers):
- Box: Middle 50% of data (interquartile range, IQR)
- Line in box: Median value
- Whiskers: Extend to 1.5× IQR or min/max values
- Small dots: Individual outliers beyond whiskers
-
Red diamonds with error bars:
- Diamond: Mean (average) value
- Error bars: Standard error of the mean (SEM = SD/√n)
-
Significance brackets (if present):
- Lines above boxes show significant pairwise differences
-
*p < 0.05,**p < 0.01,***p < 0.001
Statistical Analysis:
Non-parametric testing is used because temporal birdsong metrics often show skewed distributions and non-constant variance across development:
- Kruskal-Wallis Test: An overall test to determine if significant differences exist across the three stages (BL, Post, Rec).
- Pairwise Wilcoxon Tests: Post-hoc comparisons to identify specific differences between pairs of stages (e.g., comparing Baseline to Recovery).
- Bonferroni Correction: Applied to p-values to adjust for multiple comparisons and prevent false discoveries.
-
Significance Markers: Brackets and stars
(
*) in the plot indicate significant pairwise differences based on these adjusted p-values.
Summary of Vocal Bout Changes: - Bouts become progressively shorter and contain fewer motifs from BL to Post to Rec. - The timing between bouts remains unchanged across stages.
Exporting Bout Clips
After detecting bouts, you can export them as individual audio files
for external analysis, manual review, or machine learning workflows
using create_bout_clips().
# Export up to 50 bouts per day as WAV files
sap <- create_bout_clips(
sap,
output_format = "wav",
output_dir = "exported_bouts",
n_bouts = 50, # randomly sample up to 50 bouts per day
amp_normalize = "peak", # normalize amplitude to prevent clipping
verbose = TRUE
)The exported clips are organized into a tidy directory tree
(exported_bouts/bouts/{bird_id}/{day_post_hatch}/bout_xxx.wav)
and a companion metadata.csv is written automatically with
source timing and per-clip information.
For a full walkthrough of export arguments and additional scenarios, see the Exporting Curated Song Clips vignette.
Key Parameters for Bout Detection
Understanding and optimizing these parameters is crucial for accurate bout detection:
| Parameter | Description | Default | Optimization Tips |
|---|---|---|---|
rms_threshold |
Amplitude threshold for bout detection | 0.1 |
Lower = more sensitive (detects quieter
bouts) Higher = more selective (only loud bouts) Adjust based on recording quality and background noise |
min_duration |
Minimum bout duration (seconds) | 0.5 | Filter out brief vocalizations Typical range: 0.4-0.7s Shorter for juvenile birds, longer for adults |
gap_duration |
Minimum silence between bouts (seconds) | 0.3 | Defines bout boundaries Typical: 0.3s Longer gaps = fewer, longer bouts |
freq_range |
Bandpass filter range (kHz) | c(3, 5) | Focus on vocalization frequencies Zebra finch: c(2, 8) or c(3, 5) Adjust for species-specific frequencies |
edge_window |
Time window for edge effects (seconds) | 0.05 | Handles recording start artifacts Rarely needs adjustment |
norm_method |
RMS normalization method | “quantile” | “quantile” = robust to outliers “max” = sensitive to peaks |
summary |
Include motif-bout statistics | FALSE | Set TRUE when motif data exists Adds n_motifs, align_time, bout_gap |
balanced |
Balance samples across time points | FALSE | Use TRUE for statistical comparisons Ensures equal representation |
save_plot |
Save detection visualizations | FALSE | Set TRUE to review detection quality Plots saved to base_path/plots/bout_detection/ |
plot_percent |
Percentage of files to plot | 10 | When save_plot = TRUE 10% provides good sampling |
Tips for Longitudinal Bout Analysis
1. Optimize Parameters First
Test parameters on representative files from each time point before processing the entire dataset:
# Test on a single file
test_file <- file.path(sap$base_path, "190", sap$metadata$filename[1])
# Try different thresholds
bouts_low <- find_bout(test_file, rms_threshold = 0.05, plot = TRUE)
bouts_med <- find_bout(test_file, rms_threshold = 0.1, plot = TRUE)
bouts_high <- find_bout(test_file, rms_threshold = 0.15, plot = TRUE)
# Compare results
cat("Low threshold:", nrow(bouts_low), "bouts\n")
cat("Medium threshold:", nrow(bouts_med), "bouts\n")
cat("High threshold:", nrow(bouts_high), "bouts\n")2. Check Detection Quality
Always visualize sample detections to verify bout boundaries are accurate:
# Save plots for manual review
sap <- sap |>
find_bout(
min_duration = 0.4,
save_plot = TRUE,
plot_percent = 20 # Review 20% of files
)
# Check plots in: base_path/plots/bout_detection/3. Use Summary Statistics
Enable summary = TRUE when you have motif data to
understand bout-motif relationships:
# Requires existing motif detection
sap <- sap |>
find_motif(template_name = "d", pre_time = 0.7, lag_time = 0.5) |>
find_bout(min_duration = 0.4, summary = TRUE)
# Now you can analyze motif density in bouts
sap$bouts |>
group_by(label) |>
summarise(avg_motifs_per_bout = mean(n_motifs, na.rm = TRUE))4. Balance Sampling for Comparisons
Use balanced = TRUE when creating visualizations or
statistics for cross-timepoint comparisons:
# Balanced heatmap for fair comparison
sap |> plot_heatmap(segment_type = "bouts", balanced = TRUE)
# Balanced sampling for analysis
balanced_bouts <- select_segments(
sap$bouts,
balanced = TRUE
)5. Consider Recording Quality
Adjust parameters based on recording conditions:
-
High background noise: Increase
rms_threshold, adjustfreq_range -
Variable amplitude: Use
norm_method = "quantile" -
Short recordings: Decrease
min_durationto capture all bouts -
Continuous singing: Increase
gap_durationto separate bouts
Session Info
sessionInfo()
#> R version 4.5.3 (2026-03-11)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> loaded via a namespace (and not attached):
#> [1] digest_0.6.39 desc_1.4.3 R6_2.6.1 fastmap_1.2.0
#> [5] xfun_0.57 cachem_1.1.0 knitr_1.51 htmltools_0.5.9
#> [9] rmarkdown_2.31 lifecycle_1.0.5 cli_3.6.5 sass_0.4.10
#> [13] pkgdown_2.2.0 textshaping_1.0.5 jquerylib_0.1.4 systemfonts_1.3.2
#> [17] compiler_4.5.3 tools_4.5.3 ragg_1.5.2 evaluate_1.0.5
#> [21] bslib_0.10.0 yaml_2.3.12 jsonlite_2.0.0 rlang_1.1.7
#> [25] fs_2.0.1