Introduction
This vignette demonstrates how to construct and organize a SAP (Song Analysis Pipeline) object for longitudinal analysis of zebra finch vocalizations. SAP objects serve as the central data structure in ASAP, managing recordings across multiple developmental time points.
Prerequisites: Before reading this vignette, we recommend completing:
- Overview: ASAP 101 - Basic ASAP functions
Compatibility with SAP2011
ASAP is designed to seamlessly work with audio recordings generated
by Sound Analysis Pro 2011 (SAP2011), the widely-used
software for bird song recording and analysis. The
create_sap_object() function automatically extracts
metadata from SAP2011’s standardized filename format.
Expected WAV Filename Format
SAP2011 generates WAV files with a specific naming convention that encodes recording metadata:
{bird_id}_{timestamp}_{month}_{day}_{hour}_{minute}_{second}.wavExample filename:
S237_42674.66837050_10_31_18_33_57.wav
│ │ │ │ │ │ └── Second (57)
│ │ │ │ │ └───── Minute (33)
│ │ │ │ └──────── Hour (18)
│ │ │ └─────────── Day (31)
│ │ └────────────── Month (10 = October)
│ └───────────────────────────── Timestamp (SAP internal)
└─────────────────────────────────── Bird ID (S237)Extracted Metadata
When you create a SAP object, ASAP automatically parses filenames to extract:
| Field | Source | Example |
|---|---|---|
bird_id |
Filename prefix | “S237” |
day_post_hatch |
Subfolder name | “190” |
recording_date |
Parsed from filename | “10-31” |
recording_time |
Parsed from filename | “18:33:57” |
label |
User-provided | “Baseline” |
Organizing Your Recording Data
ASAP expects recordings to be organized in a specific folder structure, where each subfolder represents a developmental time point:
base_path/
├── 190/ # Day 190 post-hatch
│ ├── S237_42674.66837050_10_31_18_33_57.wav
│ ├── S237_42674.67577440_10_31_18_46_17.wav
│ └── ...
├── 201/ # Day 201 post-hatch
│ ├── S237_42685.1754581_11_11_0_29_14.wav
│ └── ...
└── 203/ # Day 203 post-hatch
├── S237_42687.72024667_11_13_20_0_24.wav
└── ...Note: Subfolder names are used as
day_post_hatchvalues. While typically numeric days, any consistent naming convention works (e.g., “pre”, “post”, “recovery”).
Creating a SAP Object
library(ASAP)
# Create SAP object from organized recording folders
sap <- create_sap_object(
base_path = "/path/to/recordings",
subfolders_to_include = c("190", "201", "203"),
labels = c("Baseline", "Post", "Recovery")
)
# View the structure
print(sap)
summary(sap)Example output:
SAP Object
===========
Base path: /path/to/recordings
Time points: 3 (190, 201, 203)
Labels: Baseline, Post, Recovery
Total files: 1247
- 190 (Baseline): 89 files
- 201 (Post): 704 files
- 203 (Recovery): 454 filesExploring SAP Object Contents
# View metadata extracted from filenames
head(sap$metadata)
# Check number of files per time point
table(sap$metadata$label)
# View unique bird IDs
unique(sap$metadata$bird_id)
# Visualize sample recordings
visualize_song(sap, n_samples = 4, random = TRUE)Example metadata structure:
| filename | bird_id | day_post_hatch | recording_date | recording_time | label |
|---|---|---|---|---|---|
| S237_42674…wav | S237 | 190 | 10-31 | 18:33:57 | Baseline |
| S237_42685…wav | S237 | 201 | 11-11 | 00:29:14 | Post |
| S237_42687…wav | S237 | 203 | 11-13 | 20:00:24 | Recovery |
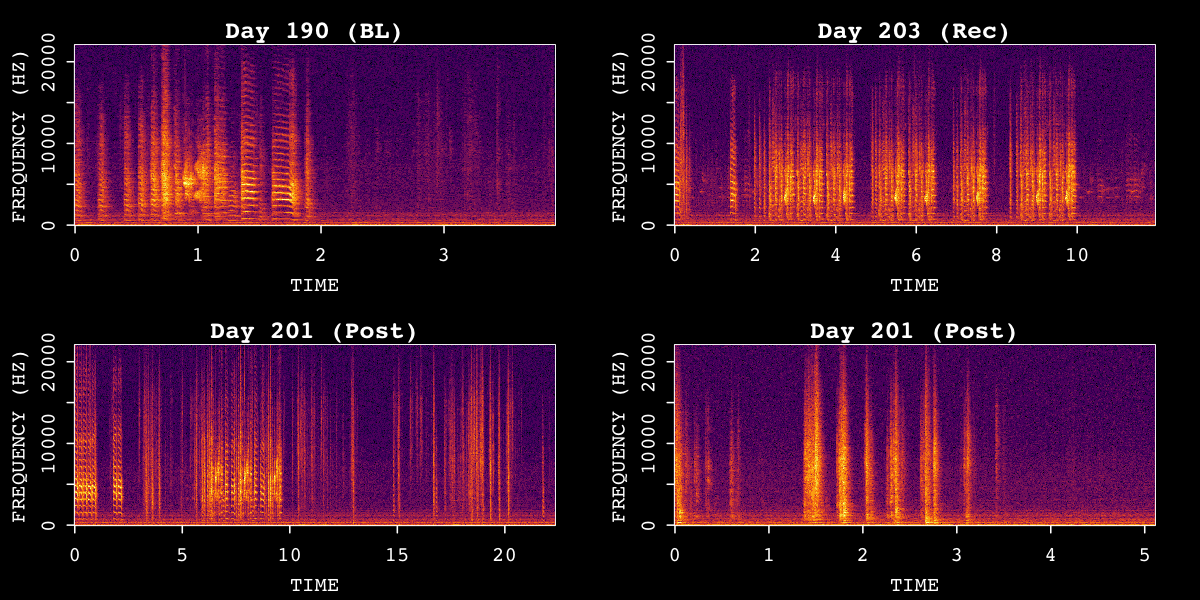
Sample spectrograms from SAP object across time points.
SAP Object Structure
A SAP object contains the following components:
| Component | Description |
|---|---|
$base_path |
Root directory path |
$metadata |
Data frame with file information (extracted from filenames) |
$templates |
Template storage (after create_template()) |
$motifs |
Detected motifs (after find_motif()) |
$bouts |
Detected bouts (after find_bout()) |
$features |
Extracted features (after analyze_spectral()) |
Working with Non-SAP2011 Recordings
ASAP was designed with flexibility in mind. If your recordings are not from SAP2011, you have several options:
Option 1: Create a Wrapper Function
You can write a simple wrapper function to rename your files to match the SAP2011 naming convention. This allows you to use all SAP object functionality directly:
# Example: Convert custom filenames to SAP format
rename_to_sap_format <- function(input_dir, output_dir, bird_id) {
files <- list.files(input_dir, pattern = "\\.wav$", full.names = TRUE)
for (f in files) {
# Extract timestamp from your custom format
# Then create SAP-compatible filename
timestamp <- as.numeric(Sys.time())
time_parts <- format(Sys.time(), "%m_%d_%H_%M_%S")
new_name <- sprintf("%s_%.5f_%s.wav", bird_id, timestamp, time_parts)
file.copy(f, file.path(output_dir, new_name))
}
}Option 2: Custom Metadata Object
For datasets that don’t follow ASAP’s naming convention—or for multi-modal experiments with synchronized neural/behavioral data—you can create a custom metadata object directly.
# List your audio files
my_files <- list.files("/path/to/recordings",
pattern = "\\.wav$",
recursive = TRUE, full.names = TRUE
)
# Create metadata with your own parsing logic
my_metadata <- data.frame(
filename = basename(my_files),
bird_id = extract_bird_id(my_files),
day_post_hatch = extract_day(my_files),
recording_date = extract_date(my_files),
recording_time = extract_time(my_files),
label = assign_labels(my_files),
# Optional: add columns for multi-modal synchronization
photometry_file = get_matched_photometry(my_files),
neural_timestamp = get_neural_sync(my_files),
stringsAsFactors = FALSE
)This approach supports integration with fiber photometry, electrophysiology, or behavioral video data. For a complete example, see: Juvenile DA Analysis - Data Processing
Next Steps
Once you have created a SAP object, proceed to:
- Longitudinal Motif Detection - Apply templates matching across all recordings
Session Info
sessionInfo()
#> R version 4.5.3 (2026-03-11)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> loaded via a namespace (and not attached):
#> [1] digest_0.6.39 desc_1.4.3 R6_2.6.1 fastmap_1.2.0
#> [5] xfun_0.57 cachem_1.1.0 knitr_1.51 htmltools_0.5.9
#> [9] rmarkdown_2.31 lifecycle_1.0.5 cli_3.6.5 sass_0.4.10
#> [13] pkgdown_2.2.0 textshaping_1.0.5 jquerylib_0.1.4 systemfonts_1.3.2
#> [17] compiler_4.5.3 tools_4.5.3 ragg_1.5.2 evaluate_1.0.5
#> [21] bslib_0.10.0 yaml_2.3.12 jsonlite_2.0.0 rlang_1.1.7
#> [25] fs_2.0.1